Scrambled Genes in Ciliates:
Introduction:
Ciliated protozoans (Phylum Ciliophora) are characterized by the presence of cilia – used for locomotion, and the presence of two
types of nuclei: a somatic nucleus – macronucleus (MAC) – which provides templates for the transcription of all genes required for
vegetative growth, and a genetic nucleus – micronucleus (MIC) – used for the exchange of meiotic products during sexual reproduction.
Following conjugation (sexual reproduction), during which haploid gametic nuclei are swapped between pairs of mating cells and a diploid zygotic nucleus formed, new MIC and MAC are generated from copies of the zygotic nucleus. DNA in the MIC remains organized in the form typical of most eukaryotes, with pairs of large chromosomes. In contrast, chromosomes in the MAC genome undergo extensive DNA fragmentation, elimination and amplification, followed by telomere addition (Prescott 1994), resulting in many small acentric chromosomes.
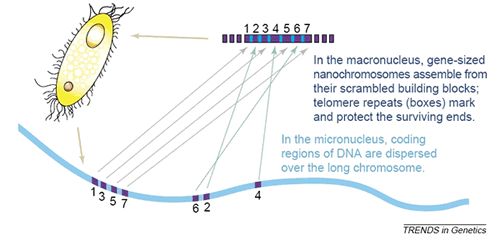
The extent of MAC genome reorganization varies greatly among ciliate species. In ciliates belonging to the class Spirotrichea the level of DNA processing in the formation of a new MAC is extraordinary: the original zygotic chromosomes are fragmented at tens of thousands of positions, 95% of the DNA complexity is lost, and the resulting chromosomes – sometimes referred to as "nanochromosomes" – are amplified to thousands of copies each (Prescott 1994). In spirotrichs, each MAC chromosome typically contains a single gene flanked at the 5' and 3' ends by very short (<150 nt) untranslated regions and telomeres (Prescott and Dizick 2000; Cavalcanti et al. 2004a). The size of these molecules ranges from 0.25 kb to 35kb, and each is present at an average of 1000 copies.
Not only is intergenic DNA lost during the conversion from MIC to MAC, but genes in the MIC are interrupted by non-coding A–T rich segments called internal eliminated segments (IESs) that must be removed. The parts of a gene separated by IESs are called macronuclear destined segments (MDSs). Some micronuclear genes have their MDSs present in a permuted order, compared to the functional gene in the macronucleus. These MDSs are rearranged during MAC formation in a deterministic way to reconstruct the functional form of the gene (Figure 1; Landweber et al., 2000; Prescott, 2000). To date, there are very little sequence data from micronuclear genes in Stichotrichs; however, in Oxytricha trifallax (also called Sterkiella histriomuscorum), three of the six genes for which MIC and MAC sequences are available are scrambled. If this is at all representative, it suggests that scrambled genes are extremely common in these organisms.
How these organisms manage to unscramble the genes is still a mystery. However, MDSs share some similar features. The last few nucleotides of an MDS are repeated in the beginning of the next consecutive MDS; these sequences are called pointers or junction sequences. Although pointers probably play a role in the unscrambling of the gene, they are not uniquely present in these two locations of the MIC locus – instead, in some cases, there are several copies of the same pointer sequence throughout the MIC gene (Landweber et al., 2000) – and thus cannot be used to precisely define the IES boundaries.
Gene Unscrambler: A program to unscramble ciliate genes.
The study of gene scrambling in ciliates is complicated by the difficulties in sequencing and annotating micronuclear genes. After both
forms of a gene are sequenced, the MDSs have to be aligned with the MAC sequences and IESs and pointer sequences must be determined based on
this alignment. Until recently there were no available programs to perform this alignment and annotation, a problem that would become more
prominent as genome data become available for Oxytricha trifallax (Doak et al. 2003; Cavalcanti et al. 2004a, b; see below).
Recently we developed a publicly available online program, Gene Unscrambler (Cavalcanti et al. 2004; http://oxytricha.princeton.edu/GeneUnscrambler.htm) developed to fill this gap. The program automatically aligns the macronuclear and micronuclear genes and gives the coordinates of MDSs and pointer sequences.
IES_MDS_Db: A database of Spirotrich genes.
A further difficulty in the study of ciliate gene rearrangements is the difficulty to annotate and retrive these genes from public databases,
like GenBank. The fields supported by these databases are not compatible with the complexity of the information in these genes. To fill this gap
we developed the IES_MDS database (Cavalcanti, Clarke and Landweber, 2004;
http://oxytricha.princeton.edu/dimorphism/database.htm), which contains all the genes in Spirotrichs for which both versions, micronuclear and
macronuclear, have been sequenced, and allows the user to search for genes of interest using several parameters.
Oxytricha trifallax pilot genome project.
Recently the Utah Genome Depot sequenced ~2000 macronuclear chromosomes of the spirotrich
Oxytricha trifallax (also called Sterkiella histriomuscorum), in a pilot genome project. We already begin to obtain interesting results
from these sequences (Doak et al 2003; Cavalcanti et al 2004a, b), and Oxytricha trifallax is now
part of the high priority list of organims for genome sequencing from the NHGRI (
National Human Genome Research Institute; Powell 2002)
Together with the Tetrahymena termophila ( http://www.ciliate.org/) and Paramecium tetraurelia (http://paramecium.cgm.cnrs-gif.fr/) genomes, we will soon have the full macronuclear genome sequence of three ciliate species, which will allow for comparative genomics analysis.
Cavalcanti A.R.O. and Landweber L.F. (2004) Gene Unscrambler for detangling scrambled genes in ciliates. Bioinformatics 20 (5): 800-802. ![]()
Cavalcanti A.R.O., Dunn D.M., Weiss R., Herrick G., Landweber L.F., Doak T.G. (2004a) Sequence features of Oxytricha trifallax (class Spirotrichea) macronuclear telomeric and subtelomeric sequences. Protist (in press).
Cavalcanti A.R.O., Stover N., Orecchia L., Doak T.G., Landweber L.F. (2004b) Coding properties of Oxytricha trifallax (Sterkiella histriomuscorum) macronuclear chromosomes: analysis of a pilot genome project. Chomosoma (in press).
Doak T.G., Cavalcanti A.R.O., Stover N., Dunn D.M., Weiss R., Herrick G., Landweber L.F. (2003) Sequencing the Oxytricha trifallax macronuclear genome: a pilot project. Trends Genet. 19 (11): 603-607. ![]()
Landweber L.F., Kuo T.C., Curtis E.A. (2000) Evolution and assembly of an extremely scrambled gene. PNAS 97: 3298–3303. ![]()
Powell K. (2002) Second round of gene sequencing goes down to the farm. Nature 419: 237.
Prescott D.M. (1994) The DNA of CIliated Protozoa. Microbiol. Rev. 58 (2): 233-267. ![]()
Prescott D.M., Dizick S.J. (2000) A unique pattern of intrastrand anomalies in base composition of the DNA in hypotrichs. Nucleic Acids Res. 28 (23): 4679-4688. ![]()
Prescott D.M. (2000) Genome gymnastics: unique modes of DNA evolution and processing in ciliates. Nat. Rev. Genet. 1: 191–198. ![]()